-
Notifications
You must be signed in to change notification settings - Fork 0
/
lecture3_expression_analysis.Rmd
897 lines (657 loc) · 28.5 KB
/
lecture3_expression_analysis.Rmd
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
326
327
328
329
330
331
332
333
334
335
336
337
338
339
340
341
342
343
344
345
346
347
348
349
350
351
352
353
354
355
356
357
358
359
360
361
362
363
364
365
366
367
368
369
370
371
372
373
374
375
376
377
378
379
380
381
382
383
384
385
386
387
388
389
390
391
392
393
394
395
396
397
398
399
400
401
402
403
404
405
406
407
408
409
410
411
412
413
414
415
416
417
418
419
420
421
422
423
424
425
426
427
428
429
430
431
432
433
434
435
436
437
438
439
440
441
442
443
444
445
446
447
448
449
450
451
452
453
454
455
456
457
458
459
460
461
462
463
464
465
466
467
468
469
470
471
472
473
474
475
476
477
478
479
480
481
482
483
484
485
486
487
488
489
490
491
492
493
494
495
496
497
498
499
500
501
502
503
504
505
506
507
508
509
510
511
512
513
514
515
516
517
518
519
520
521
522
523
524
525
526
527
528
529
530
531
532
533
534
535
536
537
538
539
540
541
542
543
544
545
546
547
548
549
550
551
552
553
554
555
556
557
558
559
560
561
562
563
564
565
566
567
568
569
570
571
572
573
574
575
576
577
578
579
580
581
582
583
584
585
586
587
588
589
590
591
592
593
594
595
596
597
598
599
600
601
602
603
604
605
606
607
608
609
610
611
612
613
614
615
616
617
618
619
620
621
622
623
624
625
626
627
628
629
630
631
632
633
634
635
636
637
638
639
640
641
642
643
644
645
646
647
648
649
650
651
652
653
654
655
656
657
658
659
660
661
662
663
664
665
666
667
668
669
670
671
672
673
674
675
676
677
678
679
680
681
682
683
684
685
686
687
688
689
690
691
692
693
694
695
696
697
698
699
700
701
702
703
704
705
706
707
708
709
710
711
712
713
714
715
716
717
718
719
720
721
722
723
724
725
726
727
728
729
730
731
732
733
734
735
736
737
738
739
740
741
742
743
744
745
746
747
748
749
750
751
752
753
754
755
756
757
758
759
760
761
762
763
764
765
766
767
768
769
770
771
772
773
774
775
776
777
778
779
780
781
782
783
784
785
786
787
788
789
790
791
792
793
794
795
796
797
798
799
800
801
802
803
804
805
806
807
808
809
810
811
812
813
814
815
816
817
818
819
820
821
822
823
824
825
826
827
828
829
830
831
832
833
834
835
836
837
838
839
840
841
842
843
844
845
846
847
848
849
850
851
852
853
854
855
856
857
858
859
860
861
862
863
864
865
866
867
868
869
870
871
872
873
874
875
876
877
878
879
880
881
882
883
884
885
886
887
888
889
890
891
892
893
894
895
896
897
---
title: "BCB420 - Computational Systems Biology"
subtitle: "Lecture 3 - Expression Data"
author: "Ruth Isserlin"
date: "`r Sys.Date()`"
output:
xaringan::moon_reader:
lib_dir: libs
css: [default, ./libs/example.css]
nature:
highlightStyle: github
highlightLines: true
highlightSpans: true
countIncrementalSlides: false
---
# Before we start
* Docker - feed back, questions?
* Takes forever to build. So now that we have tried out creating our own docker I have set up a base image you can use for your assignments
* [docker hub bcb420-base-image](https://hub.docker.com/repository/docker/risserlin/bcb420-base-image) -- will update image in the next few days because bioc is changing the way they manage their images. see [here](https://stat.ethz.ch/pipermail/bioc-devel/2020-January/016026.html)
* **Windows home** - upgrade to Windows education<br><br>
```{r eval=FALSE}
docker run -e PASSWORD=changeit --rm -p 8787:8787 risserlin/bcb420-base-image
```
---
```{r setup, include=FALSE}
options(htmltools.dir.version = FALSE)
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
if (!requireNamespace("GEOmetadb", quietly = TRUE))
BiocManager::install("GEOmetadb")
if (!requireNamespace("ggplot2", quietly = TRUE))
install.packages("ggplot2")
if (!requireNamespace("kableExtra", quietly = TRUE))
install.packages("kableExtra")
library(ggplot2)
library(kableExtra)
library(GEOmetadb)
library(GEOquery)
```
class: middle
# Overview
* <font size=6>What is Expression Analysis and what is it good for?</font><br>
* <font size=6>Types of expression data.</font><br>
* <font size=6>Where can I find expression data?</font><br>
* <font size=6>Tools for mining the data - GEOMetadb package</font><br>
* <font size=6>Delving into the different platforms available.</font><br>
* <font size=6>Trying to find a dataset of interest</font><br>
---
# Gene Expression Analysis
## Gene Expression, what is it and what is it good for?
* Genomics - genome scale analysis
* Instead of a handul of genes we can now look at 1000s, and 10000s genes. But what can we do with all those signals?
* How can we translate these copious amounts of signals into meaningful insights and conclusions?
???
When we talk about genomics we mean many different things. Ultimately it is the study of an individual genome but it has also become synonymouse with a genome-wide study.
---
## Course Goals
So, what are **we** going to be covering during this course?<br>

* **Reminder - Assignment #1 is on the course wiki. Get started!**
---
# Types of expression data
* **Microarray expression data** -
* for overview read - Bumgarner R. (2013). [Overview of DNA microarrays: types, applications, and their future. Current protocols in molecular biology](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4011503/#!po=31.4815), Chapter 22
* **Bulk RNASeq expression data** -
* Stark R, Grzelak M, Hadfield J. [RNA sequencing: the teenage years. Nat Rev
Genet](https://www.nature.com/articles/s41576-019-0150-2). 2019 Nov;20(11):631-656
* Lun A.T.L., Chen Y., Smyth G.K. (2016) [It’s DE-licious: A Recipe for Differential Expression Analyses of RNA-seq Experiments Using Quasi-Likelihood Methods in edgeR](https://link.springer.com/protocol/10.1007%2F978-1-4939-3578-9_19#citeas). In: Mathé E., Davis S. (eds) Statistical Genomics. Methods in Molecular Biology, vol 1418. Humana Press, New York, NY
* Anders, S., McCarthy, D., Chen, Y. et al. [Count-based differential expression analysis of RNA sequencing data using R and Bioconductor](https://www.nature.com/articles/nprot.2013.099). Nat Protoc 8, 1765–1786 (2013) doi:10.1038/nprot.2013.099
* **Single Cell RNAseq expression data**
* Hwang, B., Lee, J.H. & Bang, D. [Single-cell RNA sequencing technologies and bioinformatics pipelines](https://www.nature.com/articles/s12276-018-0071-8). Exp Mol Med 50, 96 (2018)
* Vieth, B., Parekh, S., Ziegenhain, C. et al. [A systematic evaluation of single cell RNA-seq analysis pipelines](https://www.nature.com/articles/s41467-019-12266-7). Nat Commun 10, 4667 (2019)
* **Protein Expression data using Mass spectrometry**
---
# Microarray expression - Basics

Bumgarner R. (2013). [Overview of DNA microarrays: types, applications, and their future. Current protocols in molecular biology](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4011503/#!po=31.4815), Chapter 22, Unit–22.1.. doi:10.1002/0471142727.mb2201s101 - Figure 1
???
See the example for processing microarray data in the prepartory material.
---
class: middle, center
# Microarray expression Pipeline
<img src="https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4011503/bin/nihms-563981-f0003.jpg" width="62%" />
Bumgarner R. (2013). [Overview of DNA microarrays: types, applications, and their future. Current protocols in molecular biology](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4011503/#!po=31.4815), Chapter 22, Unit–22.1.. doi:10.1002/0471142727.mb2201s101 - Figure 3
---
# RNASeq Analysis - Basics
<img src="https://www.annualreviews.org/na101/home/literatum/publisher/ar/journals/content/biodatasci/2019/biodatasci.2019.2.issue-1/annurev-biodatasci-072018-021255/20190704/images/medium/bd020139.f1.gif" width="120%" />
Van den Berge, K. et al [RNA Sequencing Data: Hitchhiker's Guide to Expression Analysis](https://www.annualreviews.org/doi/full/10.1146/annurev-biodatasci-072018-021255). Annu. Rev. Biomed. Data Sci. 2:139-73
???
complementary DNA is generated from the isolated mRNA
cDNA is sequenced
and then aligned to the reference genome - using software like Tophat, Bowtie
There are different aligners that generate different data types from this data.
alignments are further assembled and annotated.
RPKM is reads per kilobase per million mapped and FPKM is fragments per kilobase of transcript per million fragments mapped.
---
.center[<font size=8>** The Explosion of RNASeq analysis**</font>]
<img src="https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4494749/bin/nihms689002f1.jpg" width="85%" />
Reuter JA, Spacek DV, Snyder MP. [High-throughput sequencing technologies](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4494749/#!po=1.31579). Mol Cell. 2015 May 21;58(4):586-97. doi: 10.1016/j.molcel.2015.05.004. PMID: 26000844; PMCID: PMC4494749.
???
Colored according to company.
MiSeq - fast, personal benchtop sequencer, with run times as low as 4 hours and outputs intended for targeted sequencing and sequencing of small genomes
HiSeq 2500 - high-throughput applications
- outputs of 1 Tb in 6 days.
---
# Bulk RNASeq
* Multiple platforms (as seen on the previous slide)
* Multiple types - short read, long read, direct read.
* 95% of published RNASeq datasets comes from **short-read** illumina techonology
* Lots of different parameters that need to be taken into account during experimental design:
* number of samples
* sample preparation - oligo-dt enrichment which enrichs for the polyA tail or ribosomal RNA depletion
* read depth - the target number of sequence reads for each sample (generally 10-30 million reads per sample)
* read lengths - when using for differential expression we only need read lengths long enough to unambiguously identify a transcript. Anything more than that is extra
* single- vs paired-end reads - again with differential expression we only care about the read counts for a given transcript so optimizing read coverage with paired-end reads is extra that isn't required. (Often comes down to cost though)
---
# Multiple phases of Bulk RNASeq analysis
## Alignment and Assembly
* takes [FASTQ files](https://support.illumina.com/bulletins/2016/04/fastq-files-explained.html), which are generated by the sequencer (not quite but they are standard files), and maps them to the transcriptome.
* Tools that rely on an reference genome include:
* [TopHat](https://ccb.jhu.edu/software/tophat/index.shtml),
* [STAR](https://hbctraining.github.io/Intro-to-rnaseq-hpc-O2/lessons/03_alignment.html),
* [HISAT](http://www.ccb.jhu.edu/software/hisat/index.shtml)
* example FASTQ file:
```{r eval=FALSE}
@SIM:1:FCX:1:15:6329:1045 1:N:0:2
TCGCACTCAACGCCCTGCATATGACAAGACAGAATC
+
<>;##=><9=AAAAAAAAAA9#:<#<;<<<????#=
```
* There are also tools that don't rely on a refernce genome - often used when a reference genome is not available or it is incomplete.
---
## Quantification
* Simplicistically we are counting up the number of reads associated with each gene.
* There are many different tools that perform quantification each with different strength and weaknesses:
* [RSEM](https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-12-323) - generates transcript estimates
* [CuffLinks](https://github.com/cole-trapnell-lab/cufflinks) - generates transcript estimates
* [MMSeq](https://genomebiology.biomedcentral.com/articles/10.1186/gb-2011-12-2-r13) - generates transcript estimates
* [HTSeq](https://htseq.readthedocs.io/en/release_0.11.1/) - generates read counts
* Unfortunately there are also many measures associated with RNASeq reads. Some try to account for inherent variability caused by number of reads and length of transcripts/gene.
* RPKM/FPKM - read or fragments per kilobase million
* RPKM for individual gene = [read counts/(total number of reads in sample/1 000 000)]/ length of gene
* TPM - transcripts per million
* TPM for individual gene = [read counts/length of gene]/total number of read in sample/ 1000000)
* raw counts
* Within the community counts have come out as the preferrable metric to use when performing quantification with Bulk RNASeq
???
how and what is counted depends on the algorithm used to quantify as different methods make different assumptions about isoforms, gaps, ambiguous/unambiguous reads...
The difference between RPKM and FPKM is related the sequencing method. RPKM is single end reads where fpkm is for paired end reads.
Just cause counts are prefferred does not mean the rest is garbage...
---
## Normalization and Filtering
* As with any data set normalization and filtering help account for technical biases and sample issues.
* Some issues that arise with RNASeq data include:
* genes with low read adundances
* read depth/library size (RPKM tried to tackle this issue)
* transcript length (RPKM tired to tackle this issue)
* [Trimmed Mean of M-values](https://genomebiology.biomedcentral.com/articles/10.1186/gb-2010-11-3-r25) - TMM, originally published in 2010. Incorporated into the edgeR package
---
.center[<font size=8>**RNASeq Expression Analysis Pipeline**</font>]
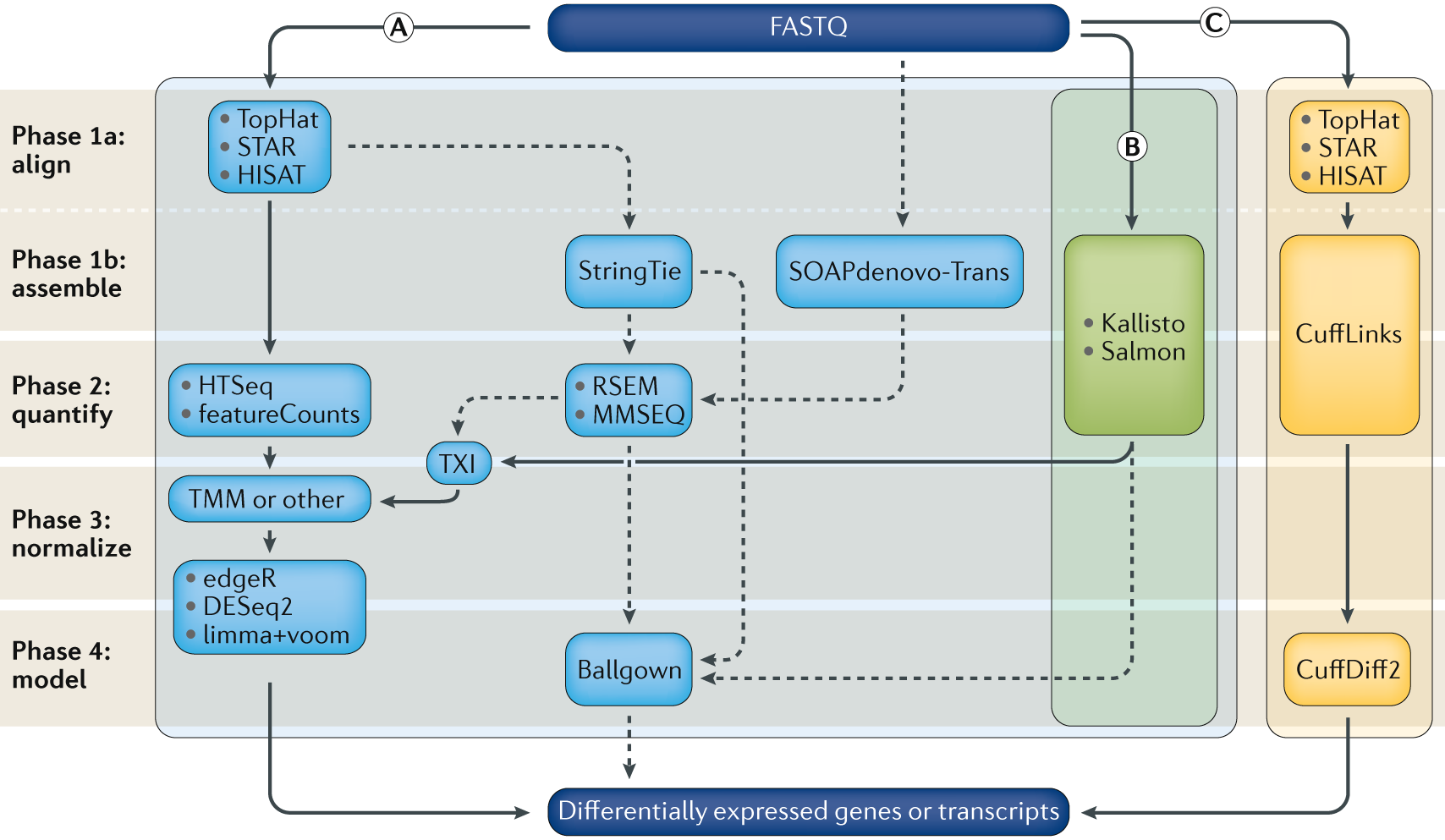
Stark, R., Grzelak, M. & Hadfield, J. [RNA sequencing: the teenage years](https://www.nature.com/articles/s41576-019-0150-2). Nat Rev Genet 20, 631–656 (2019) doi:10.1038/s41576-019-0150-2
???
Can be divided into 4 main phases Alignment/assembly, quantification, Fitering/normalization, differential expression analysis
---
# Single Cell RNASeq
* Specialized version of bulk RNASeq
* Uses a lot of the same techonologies as bulk RNASeq - **but on a much large scale**
* For data check out the [Single Cell portal - created by the broad Institute](https://singlecell.broadinstitute.org/)
* Check out these great courses that go in depth with scRNASeq
* https://github.com/broadinstitute/2019_scWorkshop
* https://scrnaseq-course.cog.sanger.ac.uk/website/index.html
---
# Protein Expression
* Protein identification and quantification via Mass spectrometry
<img src="https://els-jbs-prod-cdn.literatumonline.com/cms/attachment/cdb20d7f-724f-432d-9cc4-a1050f047496/gr1.jpg" width="85%" />
Kolker E, Higdon R, Hogan JM. [Protein identification and expression analysis
using mass spectrometry](https://www.sciencedirect.com/science/article/pii/S0966842X0600076X?via%3Dihub). Trends Microbiol. 2006 May;14(5):229-35. Epub 2006 Apr
17. Review. PubMed PMID: 16603360
---
# GEO
* Gene expression omnibus
.center[<img src="./images/geo_website.png" width="85%" />]
<font size=10>With all this data how do we figure out what to use?</font> <br><br>
???
loads and loads of databases and there are loads and loads of datasets contained in each one of these datasets.
This translates into loads of data for us to explore.
---
# GEO Datatypes
.center[<img src="https://www.ncbi.nlm.nih.gov/geo/img/geo_overview.jpg" width="85%" />]
*figure source:[GEO overview](https://www.ncbi.nlm.nih.gov/geo/info/overview.html)
???
Pop Quiz - what type of diagram is this? (Entity relationship diagram)
What do the symbols represent?
---
#GEO Datatypes - cont'd
--
**GSE - Series**<br>
* A Series record links together a group of related Samples and provides a focal point and description of the whole study.<br>
* Contains Tar archive of original raw data files, or processed sequence data files.
--
**GSM - Samples**<br>
* A Sample record describes the conditions under which an individual Sample was handled, the manipulations it underwent, and the abundance measurement of each element derived from it. <br>
* A Sample entity must reference only **one Platform** and may be included in multiple Series. <br>
* Contains Original raw data file, or processed sequence data file (for example .Cel, CPM.txt, RPKM.txt ...)<br>
---
#GEO Datatypes - cont'd
--
**GPL - Platform**<br>
* A summary description of the array or sequencer<br>
* array-based Platforms contain a data table defining the array template.<br>
* A Platform may reference many Samples that have been submitted by multiple submitters.<br>
--
**GDS - Datasets**<br>
* Curated GSEs
* same platform and calculated in the same manner (i.e background processing and normalization)
* collection of biologically and statistically comparable GEO Samples
* Not all submitted data are suitable for DataSet assembly - also there is a huge backlog so might take a long time for a series to get included in a dataset
* Works will all the GEO advanced tools.
--
**Profiles**<br>
* expression values for an individual gene across all Samples in a DataSet. <br>
---
# GEOmetadb
* a package that make querying the metadata describing microarray experiments, platforms, and datasets easier. Especially for R users.
* Based on a SQLite database that indexes all the GEO datatypes and the relationships between them. GEO datatypes including:
* GEO samples (GSM)
* GEO platforms (GPL)
* GEO data series (GSE)
* curated GEO datasets (GDS)
--
```{r message=FALSE, warning=FALSE}
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
if (!requireNamespace("GEOmetadb", quietly = TRUE))
BiocManager::install("GEOmetadb")
```
???
Pop quiz - what is the purpose of the requireNamespaces?
The function requireNamespace() is useful because it does not produce an error when a package has not been installed. It simply returns TRUE if successful or FALSE if not. Therefore one can use the following code idiom in R scripts to avoid downloading the package every time the script is called.
---
#setting up GEOmetadb
* First things first we need to do is get the meta data.
* The package doesn't download any of the expression data in GEO, just the meta data
* **Disclaimer from the GEOmetadb vignettes - "We have faithfully parsed and maintained in GEO when creating GEOmetadb. This means that limitations inherent to GEO are also inherent in GEOmetadb. We have made no attempt to curate, semantically recode, or otherwise “clean up”" GEO; to do so would require significant resources, which we do not have."**
```{r message=FALSE, warning=FALSE}
if(!file.exists('GEOmetadb.sqlite')) getSQLiteFile()
```
???
Popquiz - what is the purpose of the if statement?
---
# GEO Meta data Entity Relationship Diagram
.center[<img src="https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2639278/bin/btn520f1.jpg" width="85%" />]
Zhu Y, Davis S, Stephens R, Meltzer PS, Chen Y. [GEOmetadb: powerful
alternative search engine for the Gene Expression Omnibus](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2639278/). Bioinformatics. 2008 Dec 1;24(23):2798-800
---
# GEOmetadb DB version
```{r}
file.info('GEOmetadb.sqlite')
```
--
**Connect to our newly downloaded GEO meta data database**<br>
* When you download the database using the function getSQLiteFile, by default it places the database file into your current working directory. If you plan on using across different projects you might want to specify the download location so you don't have download everytime for a new project.
```{r}
con <- dbConnect(SQLite(),'GEOmetadb.sqlite')
```
---
# Check out the tables that are available
```{r}
geo_tables <- dbListTables(con)
geo_tables
```
---
# Look at the details of an individual table
```{r}
dbListFields(con,'gse')
```
---
# Run SQL queries
```{r}
results <- dbGetQuery(con,'select * from gpl limit 5')
knitr::kable(head(results[,1:5]), format = "html")
```
---
#Platforms
* Let's take a more detailed look at all the different platforms.
* How many are there?
--
```{r}
num_platforms <- dbGetQuery(con,'select count(*) from gpl')
```
--
```{r}
num_platforms
```
---
# Platforms - cont'd
* What other information can we get from the GPL table?
--
```{r}
dbListFields(con,'gpl')
```
---
# Platforms - cont'd
* How many different unique technologies are there?
--
```{r}
uniq_tech <- dbGetQuery(con,'select distinct technology from gpl')
nrow(uniq_tech)
```
--
We could print out the results:
```{r}
knitr::kable(uniq_tech[1:2,], format = "html") #<<
```
---
# Platforms - cont'd
Unfortunately the whole table won't fit on my slide. So instead:
```{r}
paste(uniq_tech$technology,collapse = ",")
```
---
# Platforms - cont'd
* Hom many platforms are associated with each of those platforms?
--
```{r}
num_uniq_tech <- dbGetQuery(con,'select technology,count(*) from gpl group by technology')
colnames(num_uniq_tech)[2] <- "Num_Platforms"
```
* How can we visualize this?
--
* We can print it out as a table (can't fit it into my slide)
--
* instead let's try and graph it.
```{r echo=TRUE, eval=FALSE}
plot_df <- num_uniq_tech[!is.na(num_uniq_tech$technology),]
p<-ggplot(data=plot_df, aes(technology, Num_Platforms)) +
geom_col() + coord_flip()
p
```
---
# Platforms - cont'd
```{r echo=FALSE}
plot_df <- num_uniq_tech[!is.na(num_uniq_tech$technology),]
p<-ggplot(data=plot_df, aes(technology, Num_Platforms)) +
geom_col() + coord_flip()
p
```
???
There are 18 main techonologies
the majority of them as labelled as "high throughput sequencing" or "in situ oligonucleotide" - i.e RNASequencing or microarray
---
# Platforms - cont'd
* Does this distribution change if we limit the platforms to only human
--
```{r}
num_uniq_tech_human <- dbGetQuery(con,'select technology,count(*) from gpl where organism = "human" group by technology')
colnames(num_uniq_tech_human)[2] <- "Num_Platforms"
```
--
```{r}
dim(num_uniq_tech_human)
```
Uh oh? What did I do wrong there?
---
# Platforms - cont'd
* How can I check what GEOmetadb is using for organisms? Why do I ask this? What are the possibilities?
--
```{r}
species_ids <- dbGetQuery(con,'select organism,count(*) as num_plat from gpl where organism like "%homo%" group by organism order by num_plat desc')
```
--
* Output the top 5 results:
```{r}
knitr::kable(species_ids[1:5,], format = "html")
```
???
We could use proper name, taxonomy id
---
# Platforms - cont'd
Let's get back to the original question.
* Does this distribution change if we limit the platforms to only human?
* let's use only the platforms that are defined as "Homo sapiens" only.
```{r}
num_uniq_tech_human <- dbGetQuery(con,'select technology,count(*) as num_plat from gpl where organism = "Homo sapiens" group by technology order by num_plat desc')
colnames(num_uniq_tech_human)[2] <- "Num_Platforms"
dim(num_uniq_tech_human)
```
* How would the above statement be different if I didn't care whether it was just Homo sapiens or not?
--
```{r}
num_uniq_tech_contain_human <- dbGetQuery(con,'select technology,count(*) as num_plat from gpl where organism like "%Homo sapiens%" group by technology order by num_plat desc')
colnames(num_uniq_tech_contain_human)[2] <- "Num_Platforms"
dim(num_uniq_tech_contain_human)
```
---
```{r}
knitr::kable(num_uniq_tech_human, format = "html")
```
---
```{r echo=FALSE, message=FALSE, warning=FALSE}
plot_df <- num_uniq_tech_human[
!is.na(num_uniq_tech_human$technology),]
p<-ggplot(data=plot_df, aes(technology, Num_Platforms)) +
geom_col() + coord_flip() +
ggtitle("Number of Platforms defined as only Human")
p
```
---
```{r echo=FALSE}
plot_df <- num_uniq_tech_contain_human[
!is.na(num_uniq_tech_contain_human$technology),]
p<-ggplot(data=plot_df, aes(technology, Num_Platforms)) +
geom_col() + coord_flip() +
ggtitle("Number of Platforms that contain Human")
p
```
---
* How can we compare these two sets of results better?
--
* let's merge the two sets of results that we have
* num_uniq_tech_contain_human
* num_uniq_tech_human_only
```{r}
uniq_tech_human <- rbind(data.frame(num_uniq_tech_human,
type="human only"),
data.frame(num_uniq_tech_contain_human,
type = "contains human"))
```
* what does the above dataframe look like?
--
```{r}
knitr::kable(uniq_tech_human[c(1,2,16,17),], format = "html")
```
---
class: center, middle
```{r echo=FALSE, message=FALSE, warning=FALSE}
p<-ggplot(data=uniq_tech_human, aes(technology, Num_Platforms,
fill=type)) +
geom_col(position="dodge", stat="identity") + coord_flip() +
ggtitle("Platforms that contain human vs human only ") +
theme(legend.position="bottom")
p
```
---
* This actually doesn't answer the original question.
* The original question was "Does the overall distribution change if we look for human only?"
--
* Add the data for non-organism specific query we did initially
```{r}
uniq_tech <- rbind(uniq_tech_human,
data.frame(num_uniq_tech, type = "All species"))
```
* Create the plot
```{r eval=FALSE, message=FALSE, warning=FALSE}
p<-ggplot(data=uniq_tech, aes(technology, Num_Platforms,
fill=type)) +
geom_col(position="dodge", stat="identity") +
coord_flip() +
ggtitle("Platforms that contain human vs human only ") +
theme(legend.position="bottom")
p
```
---
class: center, middle
```{r echo=FALSE, message=FALSE, warning=FALSE}
p<-ggplot(data=uniq_tech, aes(technology, Num_Platforms, fill=type)) +
geom_col(position="dodge", stat="identity") + coord_flip() + ggtitle("Platforms that contain human vs human only ") + theme(legend.position="bottom")
p
```
---
#Platforms - conclusions
* there are 18 main technology types defined for platforms.
* unfortunately they are not very specific.
* Some platforms are attributed to multiple organisms - ie. organism is not a unique field in the GEO meta data
* demonstrative exercise what we can do with the meta data available in the GEO metadb package.
---
#How to find datasets of interest?
* Let's look for datasets that are:
* RNASeq data
* recent - within the last few years
* human
* minimum number of samples
---
# GEO Meta data Entity Relationship Diagram
.center[<img src="https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2639278/bin/btn520f1.jpg" width="85%" />]
Zhu Y, Davis S, Stephens R, Meltzer PS, Chen Y. [GEOmetadb: powerful
alternative search engine for the Gene Expression Omnibus](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2639278/). Bioinformatics. 2008 Dec 1;24(23):2798-800
---
* Let's look for datasets that are:
* RNASeq data
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title",
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gpl.title LIKE '%HiSeq%' ", #<<
sep=" ")
```
--
```{r}
rs <- dbGetQuery(con,sql)
dim(rs)
```
---
* Alternately, use the technology tag
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title",
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gpl.technology = 'high-throughput sequencing' ", #<<
sep=" ")
```
--
```{r}
rs <- dbGetQuery(con,sql)
dim(rs)
```
---
* Let's look for datasets that are:
* RNASeq data
* human
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title",
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gpl.title LIKE '%HiSeq%' AND ",
" gpl.organism = 'Homo sapiens'", #<<
sep=" ")
```
--
```{r}
rs <- dbGetQuery(con,sql)
dim(rs)
```
---
* Let's look for datasets that are:
* RNASeq data
* human
* dataset from within 5 years
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title,",
" gse.submission_date",
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gse.submission_date > '2015-01-01' AND", #<<
" gpl.organism LIKE '%Homo sapiens%' AND",
" gpl.title LIKE '%HiSeq%' ",
sep=" ")
```
--
```{r}
rs <- dbGetQuery(con,sql)
dim(rs)
```
---
* Let's look for datasets that are:
* RNASeq data
* human
* dataset from within 5 years
* related to ovarian cancer
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title,",
" gse.submission_date",
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gse.submission_date > '2015-01-01' AND",
" gse.title LIKE '%ovarian cancer%' AND", #<<
" gpl.organism LIKE '%Homo sapiens%' AND",
" gpl.title LIKE '%HiSeq%' ",sep=" ")
```
--
```{r}
rs <- dbGetQuery(con,sql)
dim(rs)
```
---
* Let's look for datasets that are:
* RNASeq data
* human
* dataset from within 5 years
* related to ovarian cancer
* supplementary file is counts - unfortunately not so straightforward
```{r}
sql <- paste("SELECT DISTINCT gse.title,gse.gse, gpl.title,",
" gse.submission_date,",
" gse.supplementary_file", #<<
"FROM",
" gse JOIN gse_gpl ON gse_gpl.gse=gse.gse",
" JOIN gpl ON gse_gpl.gpl=gpl.gpl",
"WHERE",
" gse.submission_date > '2015-01-01' AND",
" gse.title LIKE '%ovarian cancer%' AND",
" gpl.organism LIKE '%Homo sapiens%' AND",
" gpl.title LIKE '%HiSeq%' ",
" ORDER BY gse.submission_date DESC",sep=" ")
```
---
* We have added the supplementary file information.
* The file name itself is determined by the submitter. What does that translate into?
--
* **A free for all**
```{r}
rs <- dbGetQuery(con,sql)
# break the file names up and just get the actual file name
unlist(lapply(rs$supplementary_file,
FUN = function(x){x <- unlist(strsplit(x,";")) ;
x <- x[grep(x,pattern="txt",ignore.case = TRUE)];
tail(unlist(strsplit(x,"/")),n=1)})) [1:10]
```
--
* Let's try and get samples that have counts data.
```{r}
rs <- dbGetQuery(con,sql)
counts_files <- rs$supplementary_file[grep(rs$supplementary_file,
pattern = "count",ignore.case = TRUE)]
series_of_interest <- rs$gse[grep(rs$supplementary_file,
pattern = "count",ignore.case = TRUE)]
shortened_filenames <- unlist(lapply(counts_files,
FUN = function(x){x <- unlist(strsplit(x,";")) ;
x <- x[grep(x,pattern="count",ignore.case = TRUE)];
tail(unlist(strsplit(x,"/")),n=1)}))
shortened_filenames
```
---
# Series of Interest
```{r echo=FALSE}
results <- cbind(series_of_interest,
rs$title[grep(rs$supplementary_file,
pattern = "count",ignore.case = TRUE)])
colnames(results)[2] <- "Series Title"
knitr::kable(results, format = "html") %>%
column_spec(1, width_max = "0.25em") %>%
column_spec(2, width_min = "30em") %>%
row_spec(6, background = "yellow")
```
---
# Get the Data
```{r}
sfiles = getGEOSuppFiles('GSE70072')
fnames = rownames(sfiles)
# there is only one supplemental file
b2 = read.delim(fnames[1],header=TRUE)
head(b2)
```
---
# Next week
<font size=8> We start processing the data ....</font>